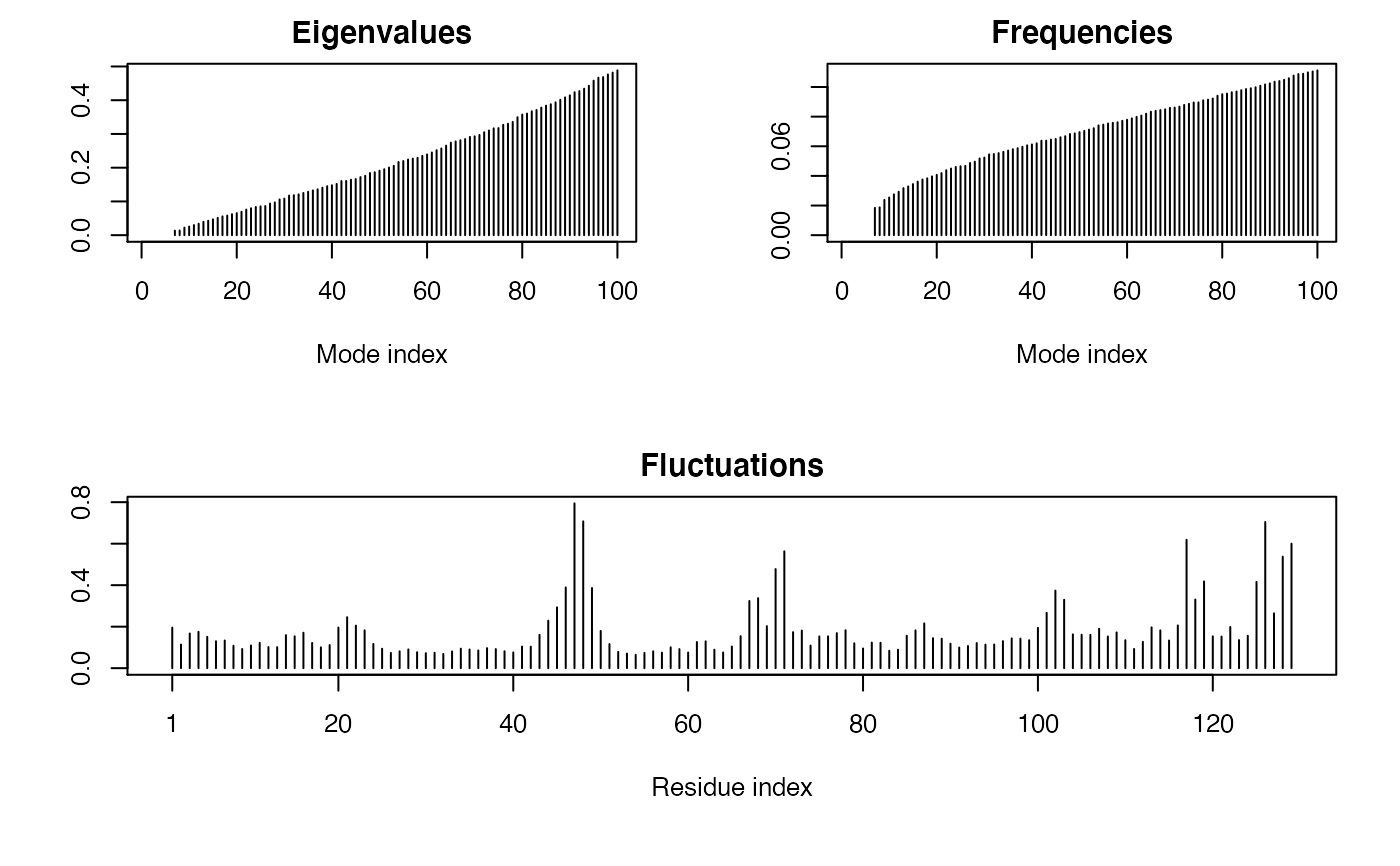
Normal Mode Analysis
nma.pdb.RdPerform elastic network model (ENM) C-alpha normal modes calculation of a protein structure.
# S3 method for pdb nma(pdb, inds = NULL, ff = 'calpha', pfc.fun = NULL, mass = TRUE, temp = 300.0, keep = NULL, hessian = NULL, outmodes = NULL, ... ) build.hessian(xyz, pfc.fun, fc.weights = NULL, pdb = NULL, ...) # S3 method for nma print(x, nmodes=6, ...)
Arguments
| pdb | an object of class |
|---|---|
| inds | atom and xyz coordinate indices obtained from
|
| ff | character string specifying the force field to use: ‘calpha’, ‘anm’, ‘pfanm’, ‘reach’, or ‘sdenm’. |
| pfc.fun | customized pair force constant (‘pfc’)
function. The provided function should take a vector of distances as
an argument to return a vector of force constants. If provided,
'pfc.fun' will override argument |
| mass | logical, if TRUE the Hessian will be mass-weighted. |
| temp | numerical, temperature for which the amplitudes for scaling the atomic displacement vectors are calculated. Set ‘temp=NULL’ to avoid scaling. |
| keep | numerical, final number of modes to be stored. Note that all subsequent analyses are limited to this subset of modes. This option is useful for very large structures and cases where memory may be limiting. |
| hessian | hessian matrix as obtained from
|
| outmodes | atom indices as obtained from |
| xyz | a numeric vector of Cartesian coordinates. |
| fc.weights | a numeric matrix of size NxN (where N is the number of calpha atoms) containg scaling factors for the pariwise force constants. See examples below. |
| x | an |
| nmodes | numeric, number of modes to be printed. |
| ... | additional arguments to |
Details
This function calculates the normal modes of a C-alpha model of a protein structure. A number of force fields are implemented all of whhich employ the elastic network model (ENM).
The ‘calpha’ force field - originally developed by Konrad Hinsen - is the recommended one for most applications. It employs a spring force constant differentiating between nearest-neighbour pairs along the backbone and all other pairs. The force constant function was parameterized by fitting to a local minimum of a crambin model using the AMBER94 force field.
See load.enmff for details of the different force fields.
By default nma.pdb will diagonalize the mass-weighted Hessian
matrix. The resulting mode vectors are moreover scaled by the thermal
fluctuation amplitudes.
The implementation under default arguments reproduces the calculation
of normal modes (VibrationalModes) in the Molecular Modeling Toolkit
(MMTK) package. To reproduce ANM modes set ff='anm',
mass=FALSE, and temp=NULL.
Value
Returns an object of class ‘nma’ with the following components:
numeric matrix with columns containing the normal mode
vectors. Mode vectors are converted to unweighted Cartesian
coordinates when mass=TRUE. Note that the 6 first trivial
eigenvectos appear in columns one to six.
numeric vector containing the vibrational
frequencies corresponding to each mode (for mass=TRUE).
numeric vector containing the force constants
corresponding to each mode (for mass=FALSE)).
numeric vector of atomic fluctuations.
numeric matrix with columns containing the raw
eigenvectors. Equals to the modes component when
mass=FALSE and temp=NULL.
numeric vector containing the raw eigenvalues.
numeric matrix of class xyz containing the
Cartesian coordinates in which the calculation was performed.
numeric vector containing the residue masses used for the mass-weighting.
numerical, temperature for which the amplitudes for scaling the atomic displacement vectors are calculated.
number of trivial modes.
number of C-alpha atoms.
the matched call.
Note
The current version provides an efficent implementation of NMA with execution time comparable to similar software (when the entire Hessian is diagonalized).
The main (speed related) bottleneck is currently the diagonalization
of the Hessian matrix which is performed with the core R function
eigen. For computing a few (5-20) approximate modes the user
can consult package ‘irlba’.
NMA is memory extensive and users should be cautions when running larger proteins (>3000 residues). Use ‘keep’ to reduce the amount of memory needed to store the final ‘nma’ object (the full 3Nx3N Hessian matrix still needs to be allocated).
We thank Edvin Fuglebakk for valuable discussions on the implementation as well as for contributing with testing.
References
Skjaerven, L. et al. (2014) BMC Bioinformatics 15, 399. Grant, B.J. et al. (2006) Bioinformatics 22, 2695--2696. Hinsen, K. et al. (2000) Chemical Physics 261, 25--37.
Author
Lars Skjaerven
See also
Examples
## Fetch stucture pdb <- read.pdb( system.file("examples/1hel.pdb", package="bio3d") ) ## Calculate normal modes modes <- nma(pdb)#> Building Hessian... Done in 0.012 seconds. #> Diagonalizing Hessian... Done in 0.076 seconds.#> #> Call: #> nma.pdb(pdb = pdb) #> #> Class: #> VibrationalModes (nma) #> #> Number of modes: #> 387 (6 trivial) #> #> Frequencies: #> Mode 7: 0.018 #> Mode 8: 0.019 #> Mode 9: 0.024 #> Mode 10: 0.025 #> Mode 11: 0.028 #> Mode 12: 0.029 #> #> + attr: modes, frequencies, force.constants, fluctuations, #> U, L, xyz, mass, temp, triv.modes, natoms, call## Visualize modes #m7 <- mktrj.nma(modes, mode=7, file="mode_7.pdb") if (FALSE) { ## Use Anisotropic Network Model modes <- nma(pdb, ff="anm", mass=FALSE, temp=NULL, cutoff=15) ## Use SSE information and SS-bonds sse <- dssp(pdb, resno=FALSE, full=TRUE) ss.bonds <- matrix(c(76,94, 64,80, 30,115, 6,127), ncol=2, byrow=TRUE) ## User defined energy function ## Note: Must take a vector of distances "my.ff" <- function(r) { ifelse( r>15, 0, 1 ) } ## Modes with a user defined energy function modes <- nma(pdb, pfc.fun=my.ff) ## A more manual approach sele <- atom.select(pdb, chain='A', elety='CA') xyz <- pdb$xyz[sele$xyz] hessian <- build.hessian(xyz, my.ff) modes <- eigen(hessian) ## Dealing with unconventional residues pdb <- read.pdb("1xj0") ## nma(pdb) #modes <- nma(pdb, mass.custom=list(CSX=121.166)) }