Overlap analysis
overlap.RdCalculate the squared overlap between sets of vectors.
overlap(modes, dv, nmodes=20)
Arguments
| modes | an object of class |
|---|---|
| dv | a displacement vector of length 3N. |
| nmodes | the number of modes in which the calculation should be based. |
Details
Squared overlap (or dot product) is used to measure the similiarity between a displacement vector (e.g. a difference vector between two conformational states) and mode vectors obtained from principal component or normal modes analysis.
By definition the cumulative sum of the overlap values equals to one.
Structure modes$U (or alternatively, the 3NxM matrix of eigenvectors)
should be of same length (3N) as dv.
Value
Returns a list with the following components:
a numeric vector of the squared dot products (overlap values)
between the (normalized) vector (dv) and each mode in mode.
a numeric vector of the cumulative squared overlap values.
References
Skjaerven, L. et al. (2011) Proteins 79, 232--243. Grant, B.J. et al. (2006) Bioinformatics 22, 2695--2696.
Author
Lars Skjaerven
See also
Examples
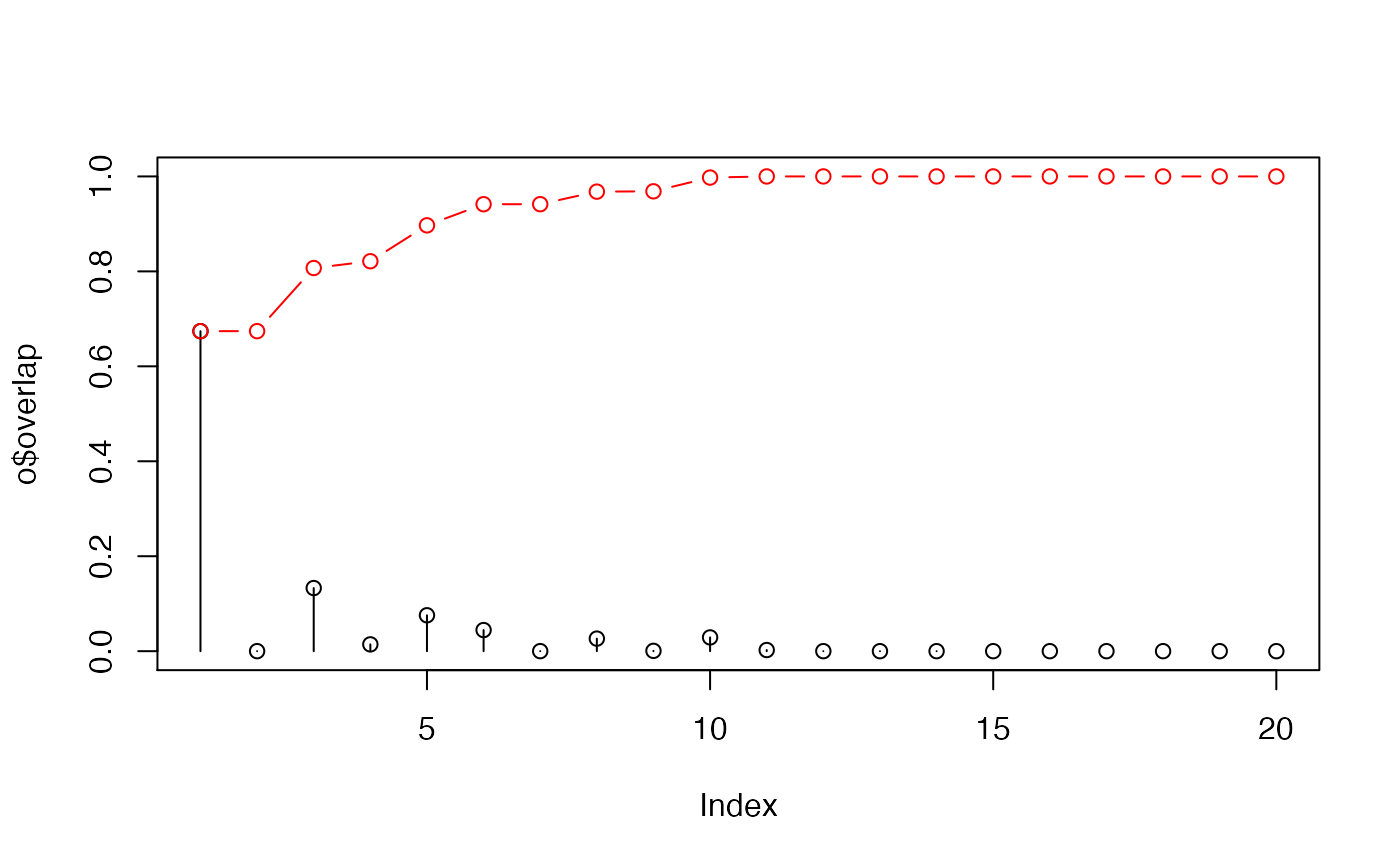
attach(kinesin) # Ignore gap containing positions ##gaps.res <- gap.inspect(pdbs$ali) gaps.pos <- gap.inspect(pdbs$xyz) #-- Do PCA pc.xray <- pca.xyz(pdbs$xyz[, gaps.pos$f.inds]) # Define a difference vector between two structural states diff.inds <- c(grep("d1v8ka", pdbs$id), grep("d1goja", pdbs$id)) dv <- difference.vector( pdbs$xyz[diff.inds,], gaps.pos$f.inds ) # Calculate the squared overlap between the PCs and the difference vector o <- overlap(pc.xray, dv) o <- overlap(pc.xray$U, dv) # Plot results plot(o$overlap, type='h', ylim=c(0,1))detach(kinesin) if (FALSE) { ## Calculate overlap from NMA pdb.a <- read.pdb("1cmk") pdb.b <- read.pdb("3dnd") ## Fetch CA coordinates sele.a <- atom.select(pdb.a, chain='E', resno=c(15:350), elety='CA') sele.b <- atom.select(pdb.b, chain='A', resno=c(1:350), elety='CA') xyz <- rbind(pdb.a$xyz[sele.a$xyz], pdb.b$xyz[sele.b$xyz]) ## Superimpose xyz[2,] <- fit.xyz(xyz[1,], xyz[2,], 1:ncol(xyz)) ## The difference between the two conformations dv <- difference.vector( xyz ) ## Calculate normal modes modes <- nma(pdb.a, inds=sele.a) # Calculate the squared overlap between the normal modes # and the difference vector o <- overlap(modes, dv) }