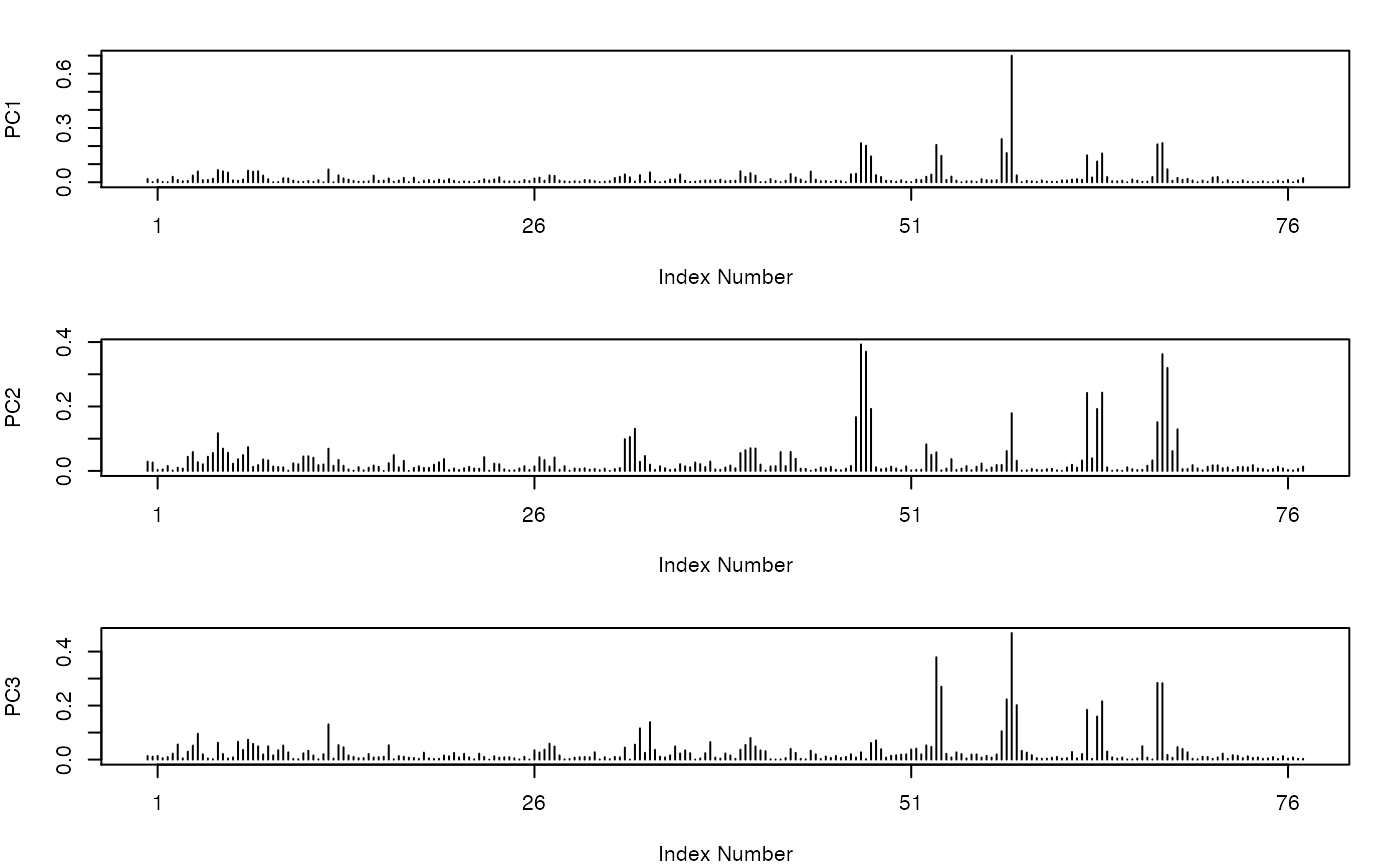
Principal Component Analysis
pca.tor.RdPerforms principal components analysis (PCA) on torsion angle data.
# S3 method for tor pca(data, ...)
Arguments
| data | numeric matrix of torsion angles with a row per structure. |
|---|---|
| ... | additional arguments passed to the method |
Value
Returns a list with the following components:
eigenvalues.
eigenvectors (i.e. the variable loadings).
scores of the supplied data on the pcs.
the standard deviations of the pcs.
the means that were subtracted.
References
Grant, B.J. et al. (2006) Bioinformatics 22, 2695--2696.
Author
Barry Grant and Karim ElSawy
See also
Examples
##-- PCA on torsion data for multiple PDBs attach(kinesin) gaps.pos <- gap.inspect(pdbs$xyz) tor <- t(apply( pdbs$xyz[, gaps.pos$f.inds], 1, torsion.xyz, atm.inc=1)) pc.tor <- pca.tor(tor[,-c(1,233,234,235)]) #plot(pc.tor) plot.pca.loadings(pc.tor)detach(kinesin) if (FALSE) { ##-- PCA on torsion data from an MD trajectory trj <- read.dcd( system.file("examples/hivp.dcd", package="bio3d") ) tor <- t(apply(trj, 1, torsion.xyz, atm.inc=1)) gaps <- gap.inspect(tor) pc.tor <- pca.tor(tor[,gaps$f.inds]) plot.pca.loadings(pc.tor) }