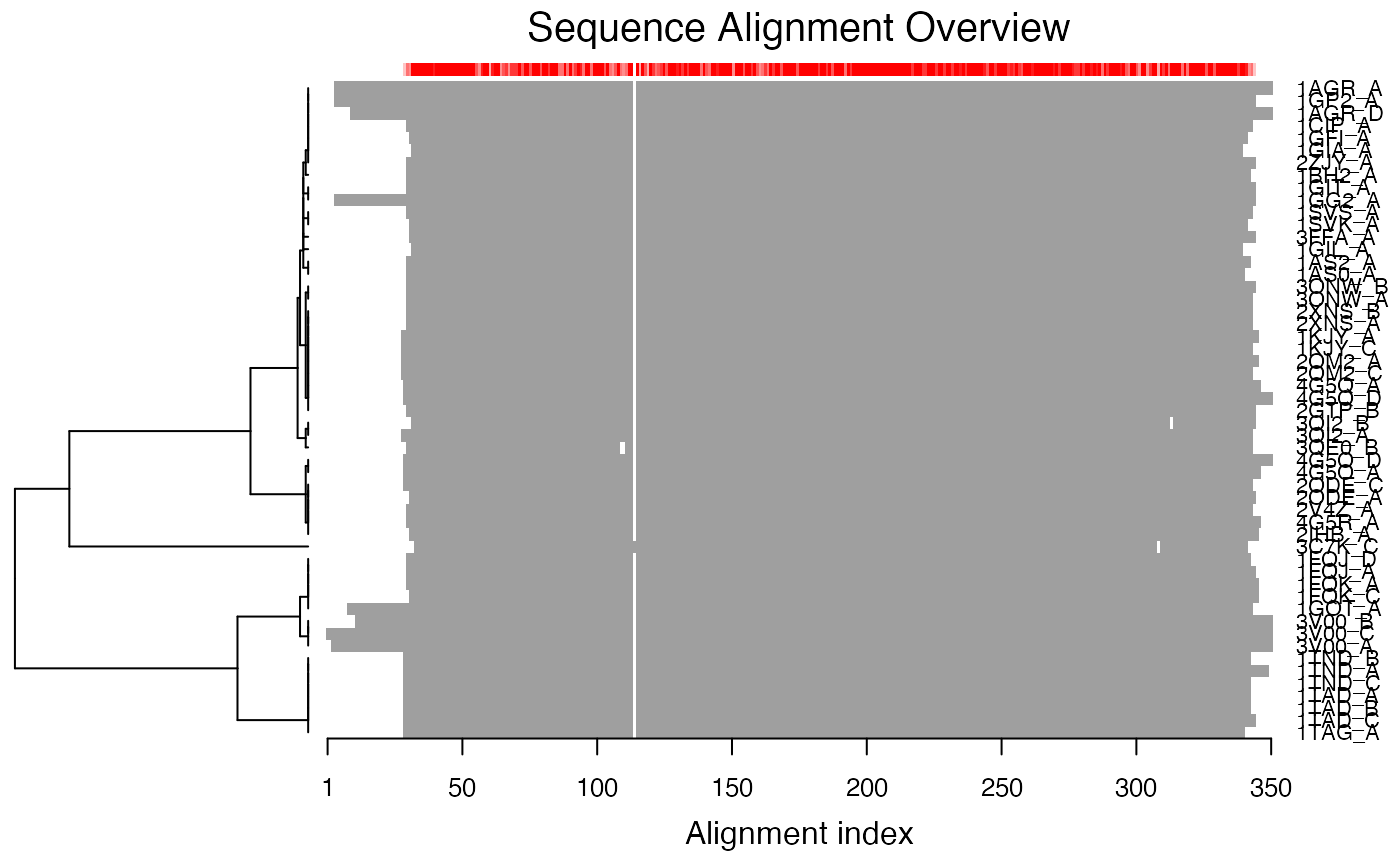
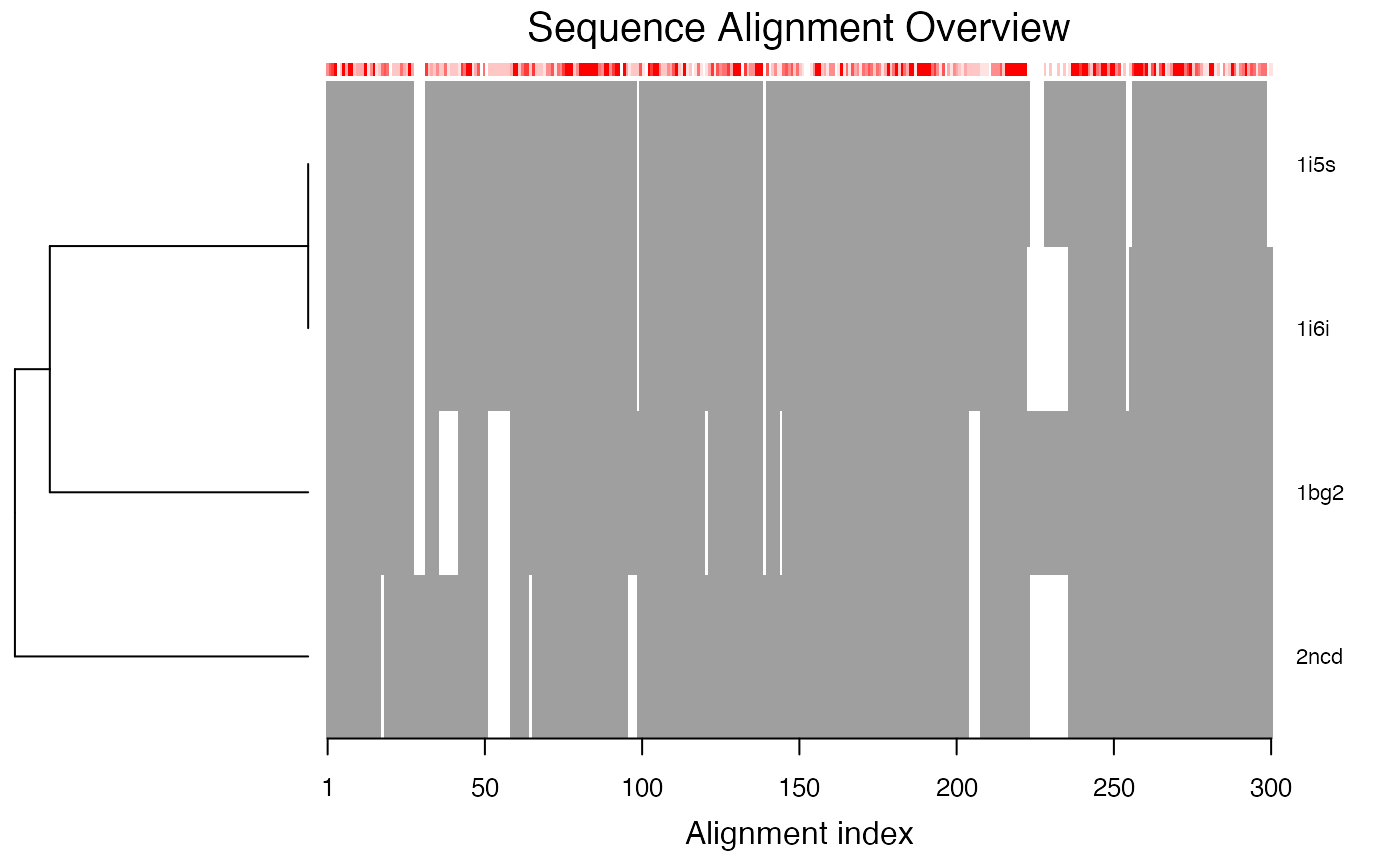
Plot a Multiple Sequence Alignment
plot.fasta.RdProduces a schematic representation of a multiple sequence alignment.
# S3 method for fasta plot(x, hc = TRUE, labels = x$id, cex.lab = 0.7, xlab = "Alignment index", main = "Sequence Alignment Overview", mar4 = 4, ...)
Arguments
| x | multiple sequence alignement of class ‘fasta’ as
obtained from |
|---|---|
| hc | logical, if TRUE plot a dendrogram on the left
side. Alternatively, an object obtained from |
| labels | labels corresponding to each row in the alignment. |
| cex.lab | scaling factor for the labels. |
| xlab | label for x-axis. |
| main | a main title for the plot. |
| mar4 | margin size for the labels. |
| ... | additional arguments passed to function |
Details
plot.fasta is a utility function for producting a schematic
representation of a multiple sequence alignment.
Value
Called for its effect.
References
Grant, B.J. et al. (2006) Bioinformatics 22, 2695--2696.
Author
Lars Skjaerven
See also
Examples
# Read alignment aln <- read.fasta(system.file("examples/kif1a.fa",package="bio3d")) ## alignment plot plot(aln, labels=basename.pdb(aln$id))detach(transducin) if (FALSE) { infile <- "http://pfam.xfam.org/family/PF00071/alignment/seed/format?format=fasta" aln <- read.fasta( infile ) plot(aln) }