Percent Identity Filter
filter.identity.RdIdentify and filter subsets of sequences at a given sequence identity cutoff.
filter.identity(aln = NULL, ide = NULL, cutoff = 0.6, verbose = TRUE, ...)
Arguments
| aln | sequence alignment list, obtained from
|
|---|---|
| ide | an optional identity matrix obtained from
|
| cutoff | a numeric identity cutoff value ranging between 0 and 1. |
| verbose | logical, if TRUE print details of the clustering process. |
| ... | additional arguments passed to and from functions. |
Details
This function performs hierarchical cluster analysis of a given sequence identity matrix ‘ide’, or the identity matrix calculated from a given alignment ‘aln’, to identify sequences that fall below a given identity cutoff value ‘cutoff’.
Value
Returns a list object with components:
indices of the sequences below the cutoff value.
an object of class "hclust", which describes the
tree produced by the clustering process.
a numeric matrix with all pairwise identity values.
References
Grant, B.J. et al. (2006) Bioinformatics 22, 2695--2696.
Author
Barry Grant
See also
Examples
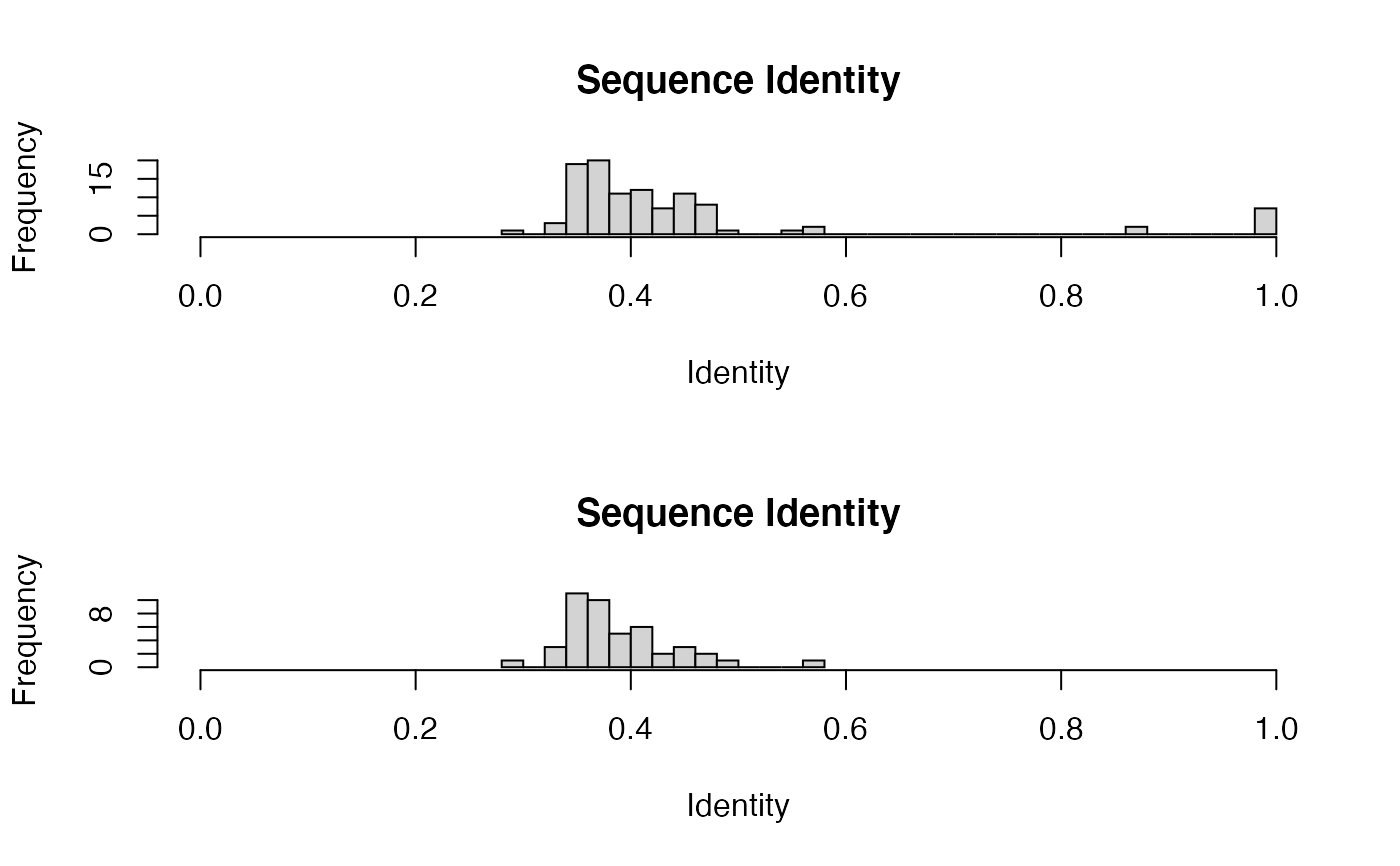
attach(kinesin) ide.mat <- seqidentity(pdbs) # Histogram of pairwise identity values op <- par(no.readonly=TRUE) par(mfrow=c(2,1)) hist(ide.mat[upper.tri(ide.mat)], breaks=30,xlim=c(0,1), main="Sequence Identity", xlab="Identity") k <- filter.identity(ide=ide.mat, cutoff=0.6)#> filter.identity(): N clusters @ cutoff = 10ide.cut <- seqidentity(pdbs$ali[k$ind,]) hist(ide.cut[upper.tri(ide.cut)], breaks=10, xlim=c(0,1), main="Sequence Identity", xlab="Identity")