Cross-Correlation for Ensemble NMA (eNMA)
Usage
"dccm"(x, ncore = NULL, na.rm=FALSE, ...)
Arguments
- x
- an object of class
enmaas obtained from functionnma.pdbs. - ncore
- number of CPU cores used to do the calculation.
ncore>1requires package ‘parallel’ installed. - na.rm
- logical, if FALSE the DCCM might containt NA values
(applies only when the
enmaobject is calculated with argument ‘rm.gaps=FALSE’). - ...
- additional arguments passed to
dccm.nma.
Description
Calculate the cross-correlation matrices from an ensemble of NMA objects.
Details
This is a wrapper function for calling dccm.nma on a collection
of ‘nma’ objects as obtained from function nma.pdbs.
See examples for more details.
Value
-
Returns a list with the following components:
- all.dccm
- an array or list containing the correlation matrices for each ‘nma’ object. An array is returned when the ‘enma’ object is calculated with ‘rm.gaps=TRUE’, and a list is used when ‘rm.gaps=FALSE’.
- avg.dccm
- a numeric matrix containing the average correlation matrix. The average is only calculated when the ‘enma’ object is calculated with ‘rm.gaps=TRUE’.
References
Wynsberghe. A.W.V, Cui, Q. Structure 14, 1647--1653. Grant, B.J. et al. (2006) Bioinformatics 22, 2695--2696.
Examples
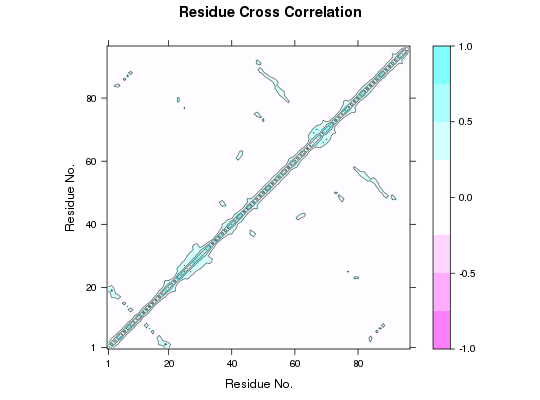
## Needs MUSCLE installed - testing excluded ## Fetch PDB files and split to chain A only PDB files ids <- c("1a70_A", "1czp_A", "1frd_A", "1fxi_A", "1iue_A", "1pfd_A") files <- get.pdb(ids, split = TRUE, path = tempdir())|======================================================================| 100%## Sequence/Structure Alignement pdbs <- pdbaln(files, outfile = tempfile())Reading PDB files: /tmp/RtmpTDihxb/split_chain/1a70_A.pdb /tmp/RtmpTDihxb/split_chain/1czp_A.pdb /tmp/RtmpTDihxb/split_chain/1frd_A.pdb /tmp/RtmpTDihxb/split_chain/1fxi_A.pdb /tmp/RtmpTDihxb/split_chain/1iue_A.pdb /tmp/RtmpTDihxb/split_chain/1pfd_A.pdb . PDB has ALT records, taking A only, rm.alt=TRUE ..... Extracting sequences pdb/seq: 1 name: /tmp/RtmpTDihxb/split_chain/1a70_A.pdb pdb/seq: 2 name: /tmp/RtmpTDihxb/split_chain/1czp_A.pdb PDB has ALT records, taking A only, rm.alt=TRUE pdb/seq: 3 name: /tmp/RtmpTDihxb/split_chain/1frd_A.pdb pdb/seq: 4 name: /tmp/RtmpTDihxb/split_chain/1fxi_A.pdb pdb/seq: 5 name: /tmp/RtmpTDihxb/split_chain/1iue_A.pdb pdb/seq: 6 name: /tmp/RtmpTDihxb/split_chain/1pfd_A.pdb## Normal mode analysis on aligned data modes <- nma(pdbs)Details of Scheduled Calculation: ... 6 input structures ... storing 282 eigenvectors for each structure ... dimension of x$U.subspace: ( 288x282x6 ) ... coordinate superposition prior to NM calculation ... aligned eigenvectors (gap containing positions removed) ... estimated memory usage of final 'eNMA' object: 3.7 Mb |======================================================================| 100%## Calculate all 6 correlation matrices cij <- dccm(modes) ## Plot correlations for first structure plot.dccm(cij$all.dccm[,,1])